Researcher Tools
Terra Platform
Terra is a cloud-native platform for biomedical researchers to access data, run analysis tools, and collaborate. The platform provides mechanisms for researchers to build, use and share packaged analytical methods and query tools, including Docker-based pipelines (e.g. for variant calling) and interactive analysis tools, including Jupyter notebooks (e.g. for visualization).
Terra is developed by the Broad Institute of MIT & Harvard and Verily.
SmartConverter Data Curation Tool
The SmartConverter tool is a set of Python and Java based custom scripts to perform data curation and update & format studies for loading into the AMP PD Knowledge Portal. The tool provides the ability to define custom data transformations and consolidation. The SmartConverter tool allows for automation of the following data curation steps: 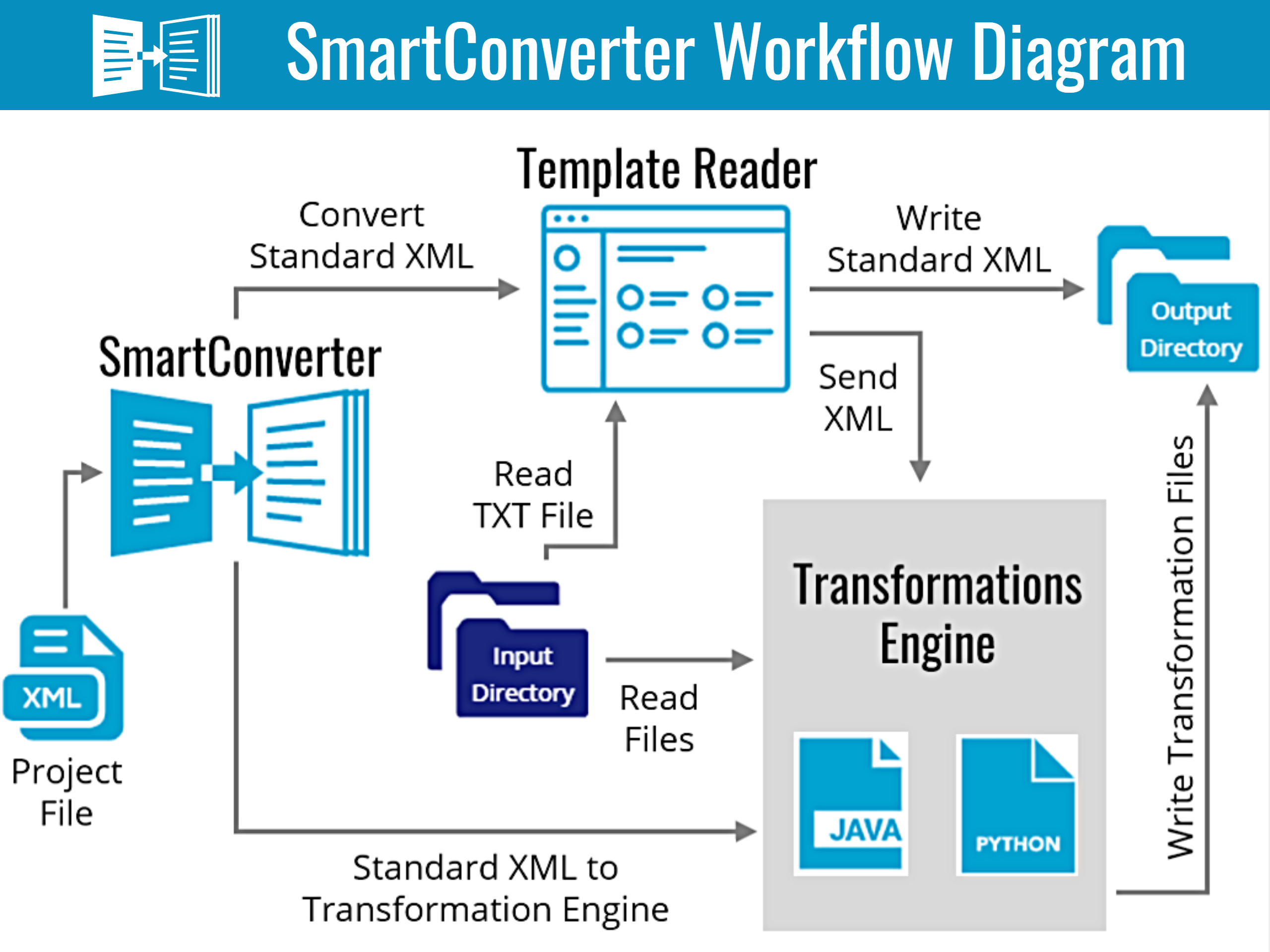
- Perform simple data transformations, logical operations, and mathematical operations
- Align data to ontologies
- Decode the data to human-readable format
- Align variable names to a harmonized dictionary
- Allows curators to store mappings/transformations for future usage
- Prepare mapping files for loading
The full dispatcher system includes:
- A dataset workflow management system
- Postgres database backend (user management, dataset status tracking and study-level metadata)
- PHP programming language
- Underlying independent scripts
- Ability to upload data from local computers or from mapped network storage files
Available Visualization & Data Explorer Tools
AMP PD offers multiple types of content and tools that are available to researchers. External to the Portal and Terra are “explorer & visualization tools,” that are developed as supplemental resources to Terra and can be used as stand-alone applications for sophisticated analysis. Researchers can use these tools to analyze AMP PD clinical, transcriptomics, and genomics data and/or in-conjunction with disparate datasets.
These tools make it easier for researchers to run canned queries and visualize results in a targeted fashion. Access to these external tools requires approval, authentication, and user login.
Terra Components
The Terra Data Explorer provides the following:
A UI that enables researchers to explore a dataset, understand its contents, and build custom cohorts for deeper analysis in Jupyter notebooks.
Terra Toolkits provides syntax editors, visualization tools, and helps you manage your workflows for a better user experience.
The Terra Interactive Analysis Service manages the loading of Jupyter notebooks and allocating backend compute services for running large computations.
Notebooks are automatically stored in a Cloud Storage bucket associated with a Workspace.
Terra Workspaces allow you to access and organize data and analysis tools, and collaborate with other researchers in Terra.
The Terra Job Manager provides a user interface for monitoring data processing workflows run by Cromwell.
Additional Terra Platform Resources:
Available & Coming Soon Tools

RNA-Seq Data Explorer
The RNA-Seq Data Explorer App provides access to gene-level RNA-seq data, quality metrics, and integration with previous DNA-based studies. Users can explore cohorts & quality, browse genomes, and perform gene-level plotting.

Bravo Variant Tool
Shows chromosome locations, VEP functional annotations, and allele frequencies. The sequencing analysis pipeline consists of two major processes: Data harmonization from BAM files and Joint variant discovery & genotype calling across studies.



