This repository is under review for potential modification in compliance with Administration directives.
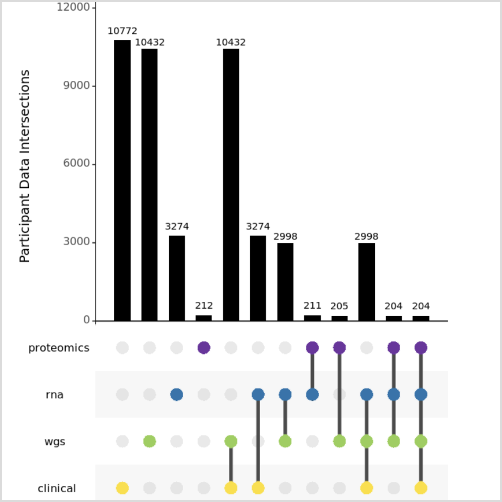
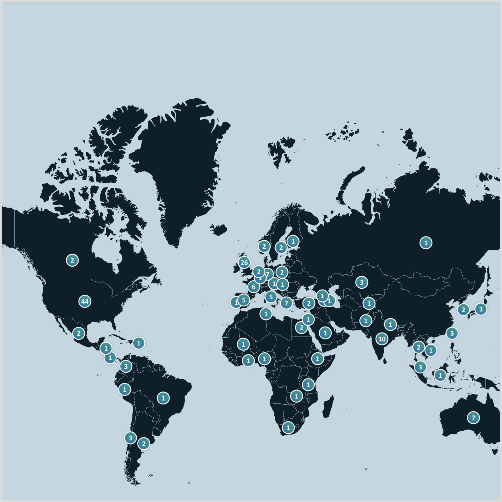
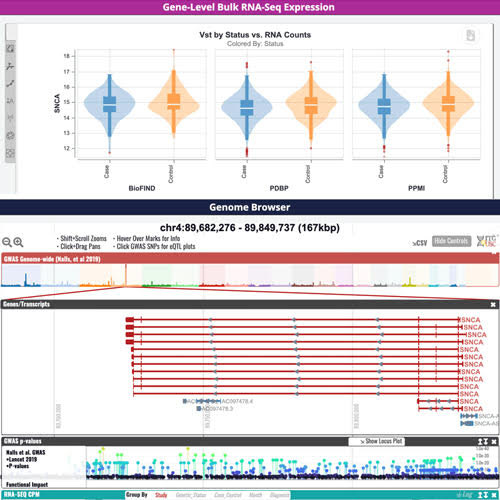
The AMP PD Knowledge Portal was developed to host and share resources related to Parkinson’s disease research and remains fully operational. We continue to maintain and accept Parkinson’s disease and related disorders data and resources throughout this review process.